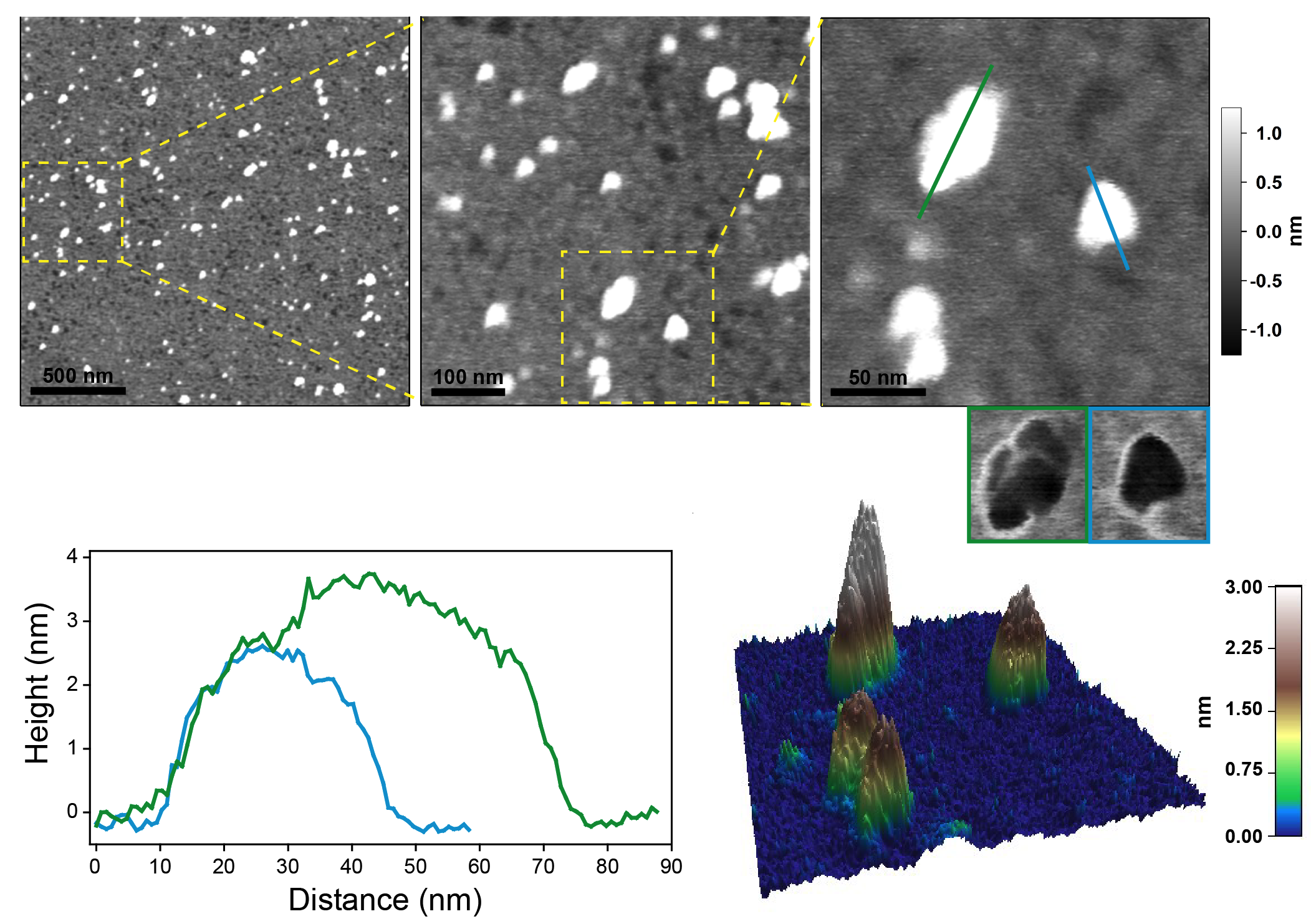
Figure: Human Arc protein, visualized via atomic force microscopy (AFM) by Anne Baumann.
In May 2011, a few months into my PhD, I took a Molecular and Computational Biology (MCB) Research School course called Recombinant proteins: Expression, Purification & Interaction Studies. Being the new, ambitious PhD candidate that I was, I started to discuss my project with fellow student, Helene Bustad Johannessen. It just so happened that her protein of interest was a proposed interaction partner with my protein. So, we made plans to test it out in the coming months. Though that specific project didn’t pan out, it was the start of a fruitful collaboration between my group (Prof. Clive Bramham) and Helene’s group (Prof. Aurora Martinez), which eventually led to last week’s publication in Biochemical Journal.
My PhD was on a protein called Arc, which is critical for long-term memory formation. Despite significant interest in Arc, nothing was known about its structure. Helene and I began by investigating Arc secondary structure with circular dichroism (CD). Anne Baumann and I learned more about Arc stability with dynamic light scattering (DLS), while Marte Innselset Flydal studied Arc stability with differential scanning calorimetry (DSC). Anne went on to do some important surface plasmon resonance (SPR) and CD experiments to show that recombinant Arc binds to a known binding partner, PS1. We also had excellent collaborators in France and Spain that added crucial parts to the paper.
Our paper is titled, Arc is a flexible modular protein capable of reversible self-oligomerization, and is the first biochemical and biophysical analysis of Arc structure and stability. We are excited about the follow-up projects on Arc structure with continued collaborations within the Jebsen Center!
Check out the paper (soon to be open access)
By Craig Myrum